 
We show that for the QM9 small molecule dataset, the best of both methods can be combined by using the conformations generated by the deep generative graph neural network as an initialization to the force field method. We also show that our approach is computationally faster than force field methods.Ī disadvantage of our model is that in general for a given molecule, the best conformation generated by our model lies further away from the reference conformation compared to the best conformation generated by force field methods. Despite having lower variance, we show that our method does not generate geometrically similar conformations. the variance of the RMSD between generated and reference conformations is lower for our method. We show that conformations generated by our model are on average far more likely to be close to the reference conformation compared to those generated by conventional force field methods i.e. We evaluate and compare our method with conventional molecular force field methods on three databases of small molecules by calculating the root-mean-square deviation (RMSD) between generated and reference conformations. This is done by maximizing the likelihood of the reference conformations of the molecules in the dataset. In this paper, we propose a deep generative graph neural network that learns the energy function from data in an end-to-end fashion by generating molecular conformations that are energetically favorable and more likely to be observed experimentally 4. Although this approach has been commonly used to generate a geometrically diverse set of conformations with certain conformations being similar to the lowest-energy conformations, it has been shown that molecule force field energy functions are often a crude approximation of actual molecular energy 3. 
The minimum of this energy function corresponds to the molecule’s most stable configuration. This hand-designed energy function yields an approximation of the molecule’s true potential energy observed in nature based on the molecule’s atoms, bonds and coordinates. Typically this problem is approached by using a force field energy function to calculate a molecule’s energy, and then minimizing this energy with respect to the molecule’s coordinates. However, these methods are typically time-consuming and costly.Ī number of computational methods have been developed for conformation generation over the past few decades 2. Conformations can be determined in a physical setting using instrumental techniques such as X-ray crystallography as well as using experimental techniques. Conformation generation is also a vital part of applications such as generating 3-D quantitative structure-activity relationships (QSAR), structure-based virtual screening and pharmacophore modeling 2. The task, known as conformation generation, of predicting possible valid coordinates of a molecule, is important for determining a molecule’s chemical and physical properties 1. The three-dimensional (3-D) coordinates of atoms in a molecule are commonly referred to as the molecule’s geometry or conformation. On one of the evaluated datasets we show that this combination allows us to combine the best of both methods, yielding generated conformations that are on average close to reference conformations with some very similar to reference conformations. We also show that our method can be used to provide initial coordinates for conventional force field methods. Our method maintains geometrical diversity by generating conformations that are not too similar to each other, and is also computationally faster. 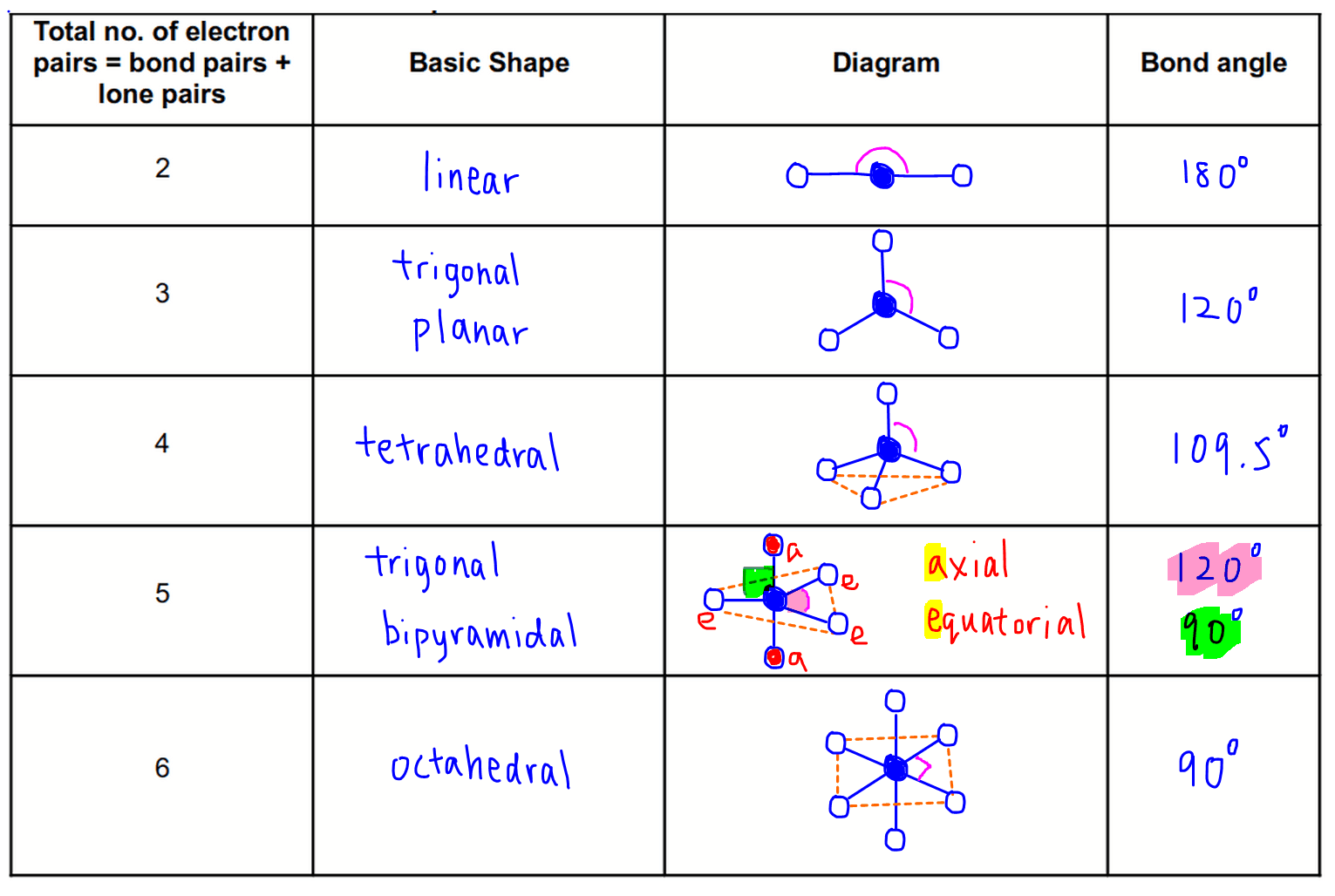
On three large-scale datasets containing small molecules, we show that our method generates a set of conformations that on average is far more likely to be close to the corresponding reference conformations than are those obtained from conventional force field methods. In this paper, we propose a conditional deep generative graph neural network that learns an energy function by directly learning to generate molecular conformations that are energetically favorable and more likely to be observed experimentally in data-driven manner. They generate geometrically diverse sets of conformations, some of which are very similar to the lowest-energy conformations and others of which are very different. Conventional conformation generation methods minimize hand-designed molecular force field energy functions that are often not well correlated with the true energy function of a molecule observed in nature. 
These structures can generally be predicted, when A is a nonmetal, using the "valence-shell electron-pair repulsion model (VSEPR) discussed in the next section.A molecule’s geometry, also known as conformation, is one of a molecule’s most important properties, determining the reactions it participates in, the bonds it forms, and the interactions it has with other molecules. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
